How Do Viruses Mutate and What it Means for a Vaccine?

As SARS-CoV-2 spreads around the globe, it is mutating, in other words it is acquiring genetic changes.
While the idea of “viral mutation” may sound concerning, it’s important to understand that many of these mutations are minor, and don’t have an overall impact on how fast a virus spreads or potentially how severe a viral infection might be. In fact, some mutations could make the virus less infectious.
Much of our knowledge of how viruses change to escape natural or vaccine-elicited immunity comes from observing the influenza virus and constantly updating influenza vaccines. Influenza viruses change in two main ways, antigenic drift and antigenic shift.
A comparison of the similarities and differences between coronaviruses and flu viruses may help us understand how those similarities and differences could impact potential COVID-19 vaccines.
Antigenic Drift
As a virus replicates, its genes undergo random “copying errors” (i.e. genetic mutations). Over time, these genetic copying errors can, among other changes to the virus, lead to alterations in the virus’ surface proteins or antigens.1
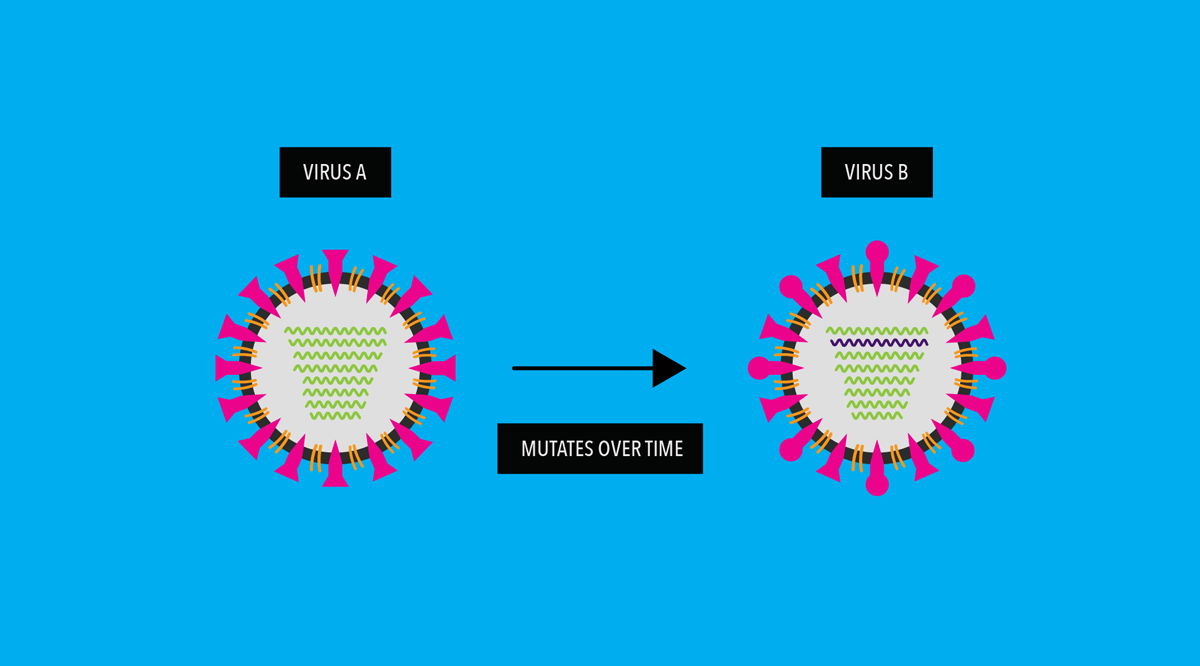
Our immune system uses these antigens to recognize and fight the virus. So, what happens if a virus mutates to evade our immune system?
In influenza viruses, genetic mutations accumulate and cause its antigens to “drift”— meaning the surface of the mutated virus looks different than the original virus.
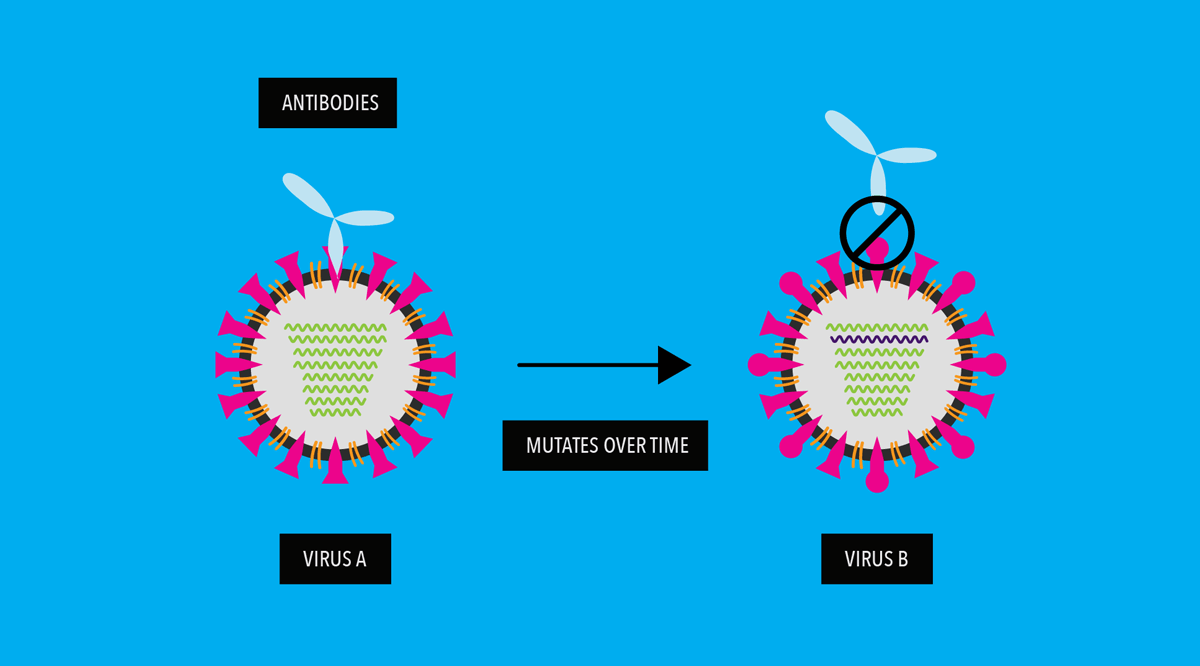
When influenza virus drifts enough, vaccines against old strains of the virus and immunity from previous influenza virus infections no longer work against the new, drifted strains. A person then becomes vulnerable to the newer, mutated flu viruses.
Antigenic drift is one of the main reasons why the flu vaccine must be reviewed and updated each year, to keep up with the influenza virus as it changes.
Could that also happen with SARS-CoV-2?
From what has been observed thus far regarding the genetic evolution of SARS-CoV-2, it appears that the virus is mutating relatively slowly as compared to other RNA viruses. Scientists think this is due to its ability to “proofread” newly made RNA copies.2 This proofreading function does not exist in most other RNA viruses, including influenza. Studies to date estimate that the novel coronavirus mutates at a rate approximately four times slower than the influenza virus, also known as the seasonal flu virus. Although SARS-CoV-2 is mutating, thus far, it does not seem to be drifting antigenically. It should be noted, however, that SARS-CoV-2 is a newly discovered virus infecting humans. There are still many unknowns, and our understanding of the SARS-CoV-2 virus continues to grow
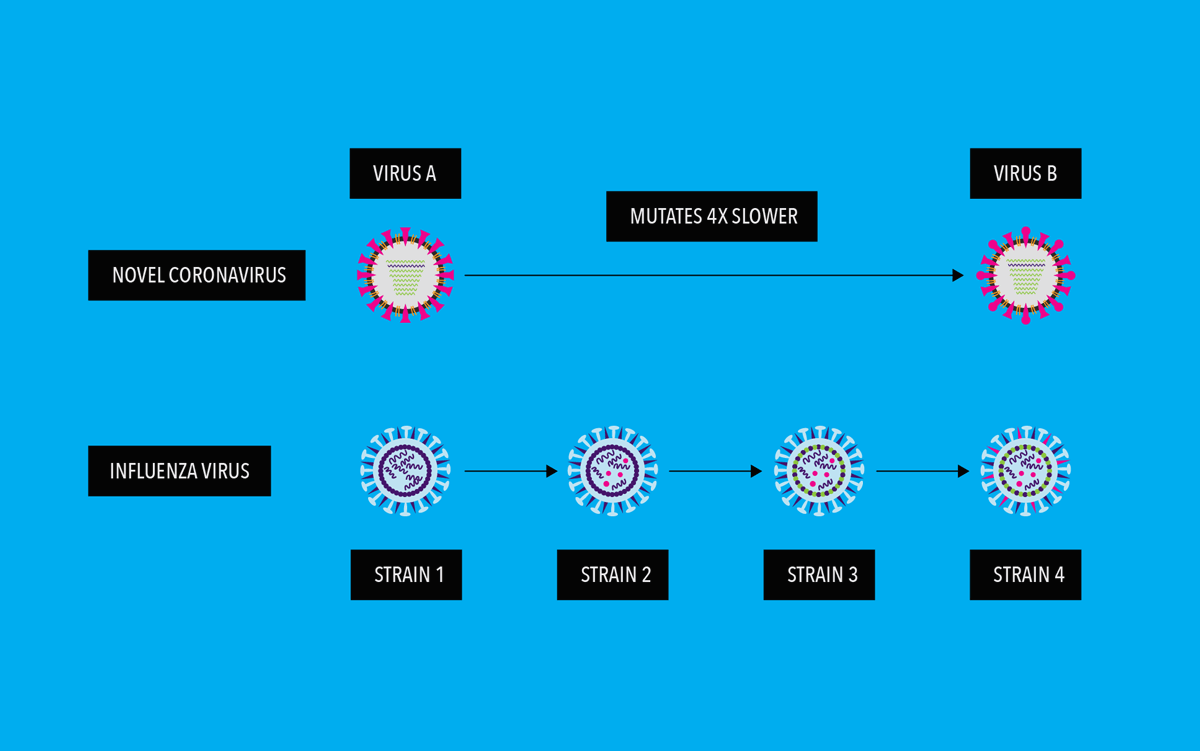
This relatively slow mutation rate for SARS-CoV-2 make us hopeful that investigational SARS-CoV-2 vaccine candidates will potentially have one less hurdle toward offering the ability to provide protection over a longer period of time.
Antigenic Shift
Influenza viruses undergo antigenic shift, an abrupt, major change in the virus’s antigens that happens less frequently than antigenic drift.3
It occurs when two different, but related, influenza virus strains infect a host cell at the same time. Because influenza virus genomes are formed by 8 separate pieces of RNA (called “genome segments”), sometimes these viruses can “mate,” in a process called, “reassortment.” During reassortment, two influenza viruses’ genome segments can combine to make a new strain of influenza virus.
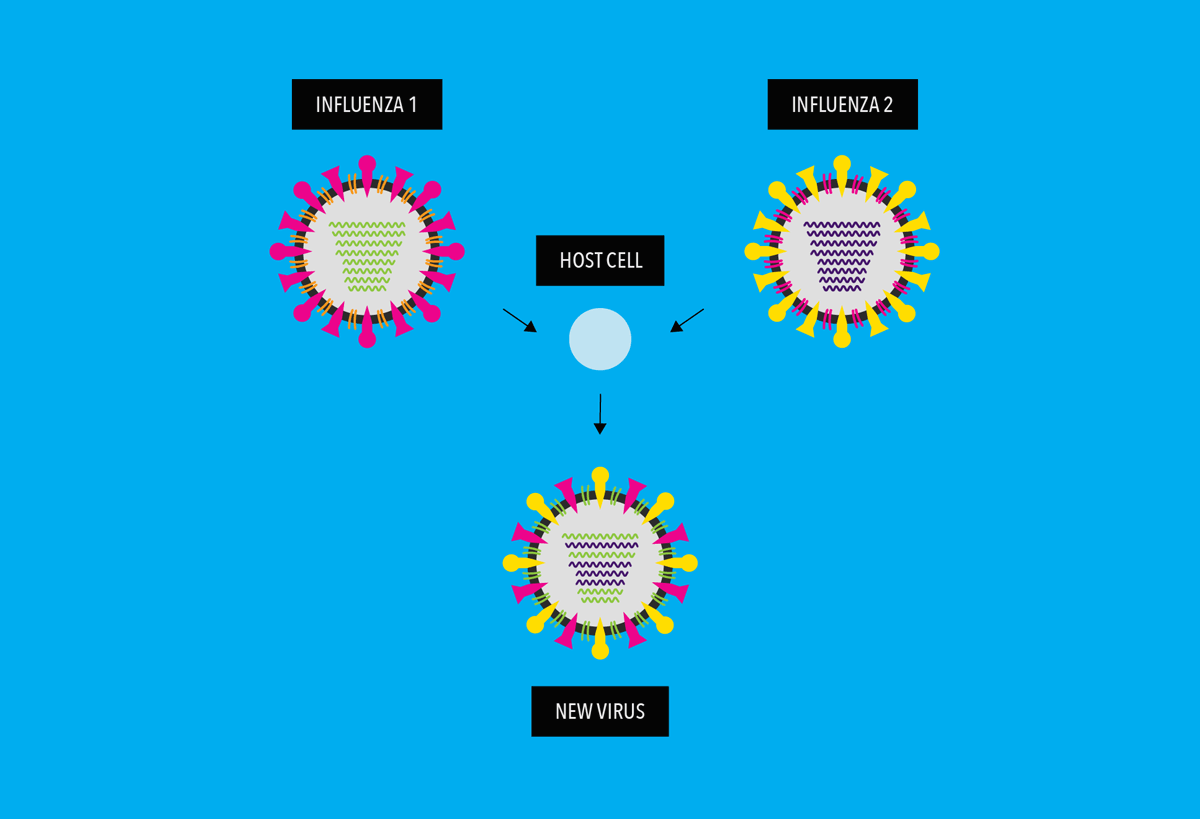
Reassortment results is a new subtype of virus, with antigens that are a mixture of the original strains.
When a shift happens, most people have little or no immunity against the resulting new virus (as shown by the “x” marks below). Viruses emerging as a result of antigenic shift are the ones most likely to cause pandemics.
Coronaviruses do not have segmented genomes and cannot reassort. Instead, the coronavirus genome is made of a single, very long piece of RNA. However, when two coronaviruses infect the same cell, they can recombine, which is different than reassortment. In recombination, a new single RNA genome is stitched together from pieces of the two “parental” coronaviruses genomes. It’s not as efficient as reassortment, but scientists believe that coronaviruses have recombined in nature.
When this happens, scientists identify the resulting virus as a “novel coronavirus.” The generation of a novel coronavirus, although occurring by a different mechanism than antigenic shift in influenza viruses, can have a similar consequence, with pandemic spread.
Alternatively, pandemic influenza viruses can sometimes arise, not by reassortment but by “zoonosis,” when an influenza virus that infects other animals, often birds or pigs, makes the leap into humans and starts to spread.
This happens with coronaviruses, too, with new human coronaviruses,4 or genes of new human coronaviruses, coming from ancestral coronavirus es that have infected other animals, such as bats, camels or pangolins.
To date, we have seen coronaviruses act like influenza viruses in generating outbreaks and now a pandemic from processes of recombination and zoonosis generating novel human coronaviruses (resembling the antigenic shift and zoonotic origin of new human influenza virus subtypes).
Thus far, we have seen human coronaviruses mutate but not undergo antigenic drift. This is good news for coronavirus vaccines. Nevertheless, given the similarities between the behavior of influenza viruses and coronaviruses, there is ample reason to remain vigilant for the possibility of future antigenic changes in SARS-CoV-2 and to be prepared to alter a potential COVID-19 vaccine, if necessary.
References
1 CDC: How Flu Viruses Can Change: “Drift” and “Shift.”
2 “Seven months later, what we know about COVID-19 — and the pressing questions that remain.” STAT. Aug. 17, 2020. https://www.statnews.com/2020/08/17/what-we-now-know-about-covid19-and-what-questions-remain-to-be-answered/
3 CDC: How Flu Virsues Can Change: “Drift” and “Shift” https://www.cdc.gov/flu/about/viruses/change.htm
4 “Coronavirus: Where do new viruses come from?” The Conversation. April 14, 2020. https://theconversation.com/coronavirus-where-do-new-viruses-come-from-136105